Supersensing for Superintelligence
How quantum sensing can improve materials/biology AI models
The current AI boom is underpinned by scaling laws. These show that model performance, quantified by loss functions like cross-entropy, gets better as you scale up compute, model capacity, and dataset size. As you can see from these plots in the original scaling laws paper, the test loss improves as you scale up these parameters with a power-law dependence.
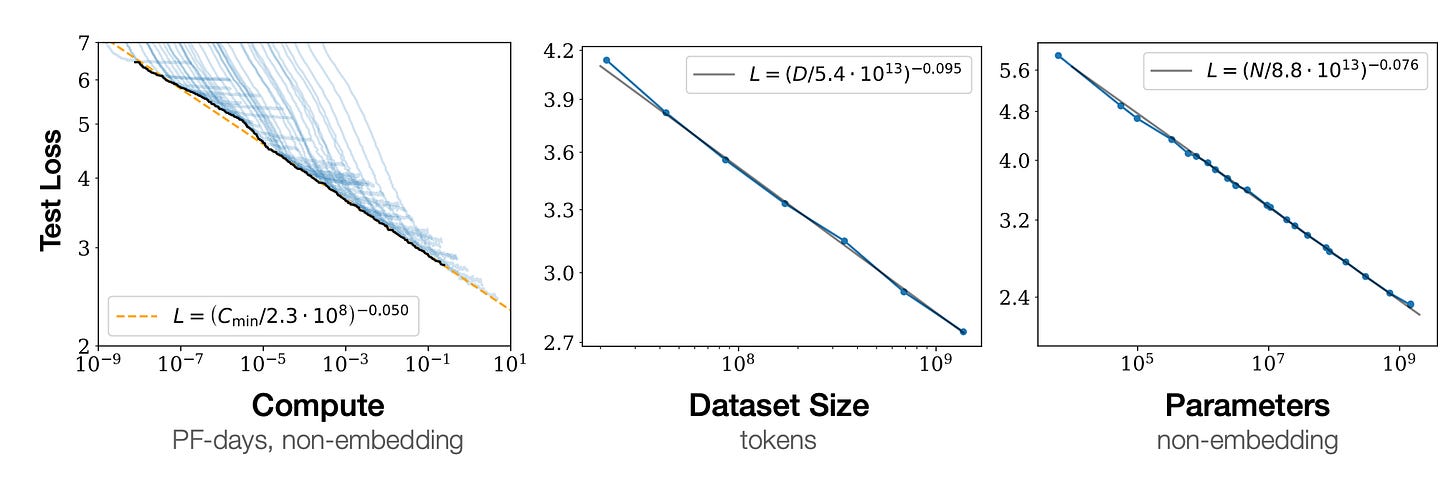
These scaling laws are responsible for the explosion in AI datacentre capex, which is now approaching trillions of dollars. More compute allows scaling models up to leviathan sizes, but a key conclusion of the scaling laws thesis is that these scalings only work when none of the three factors — compute, data, model size — is a limiting bottleneck. Later work by Google showed that LLMs were undertrained because of a “focus on scaling language models whilst keeping the amount of training data constant”. In the limit of abundant compute, data becomes the critical bottleneck.
So where can we get new, high-quality data that hasn’t already been fed to the silicon leviathans?
The answer is highly domain-dependent, but I want to focus on two areas where the opportunity is especially compelling: materials and biology, where many relevant degrees of freedom happen to be quantum mechanical. It is here that quantum sensors today and quantum computers in the future will unlock new streams of data.
As we will see, most models in these domains are trained on real, static data, i.e. snapshots of a molecule at one point in time, or synthetic dynamic data produced from good but approximate simulation methods. To build the models of the future, it’s important to get real dynamic data, which we can do with quantum sensors today, and high-quality synthetic data, for which a quantum computer is the best tool.
To gain information about the world, we need to make a measurement using a sensor. Quantum sensors are specialised to make measurements of atoms/molecules because they take into account the counterintuitive laws of quantum mechanics that govern physics at these scales. Crucially, many of these sensors already exist, meaning that these new data sources are available right now.
A great example with implications in both materials and biology today is sensitive magnetic field sensing. Sensitive magnetometry can give us information not only about the electronic and magnetic properties of materials, but also things like signal pathways in the brain, and these are invaluable sources of data to train models on.
Quantum Sensing for Materials
Materials science is a field that has attracted lots of attention for AI acceleration. DeepMind, with its GNOME model, and Periodic Labs are examples of companies doing great work in this space. However, many public materials datasets like the Materials Project and GNOME’s own dataset are missing the type of data that tells you about the dynamics of atomic systems. Luckily for us, quantum sensors are great at collecting exactly this type of data.
A workhorse of materials science, and also one of the most exquisitely precise sensors ever developed, is the Superconducting Quantum Interference Device, or SQUID. At a basic level, a SQUID consists of a loop of superconducting metal interrupted at two points by thin insulating barriers known as Josephson junctions.
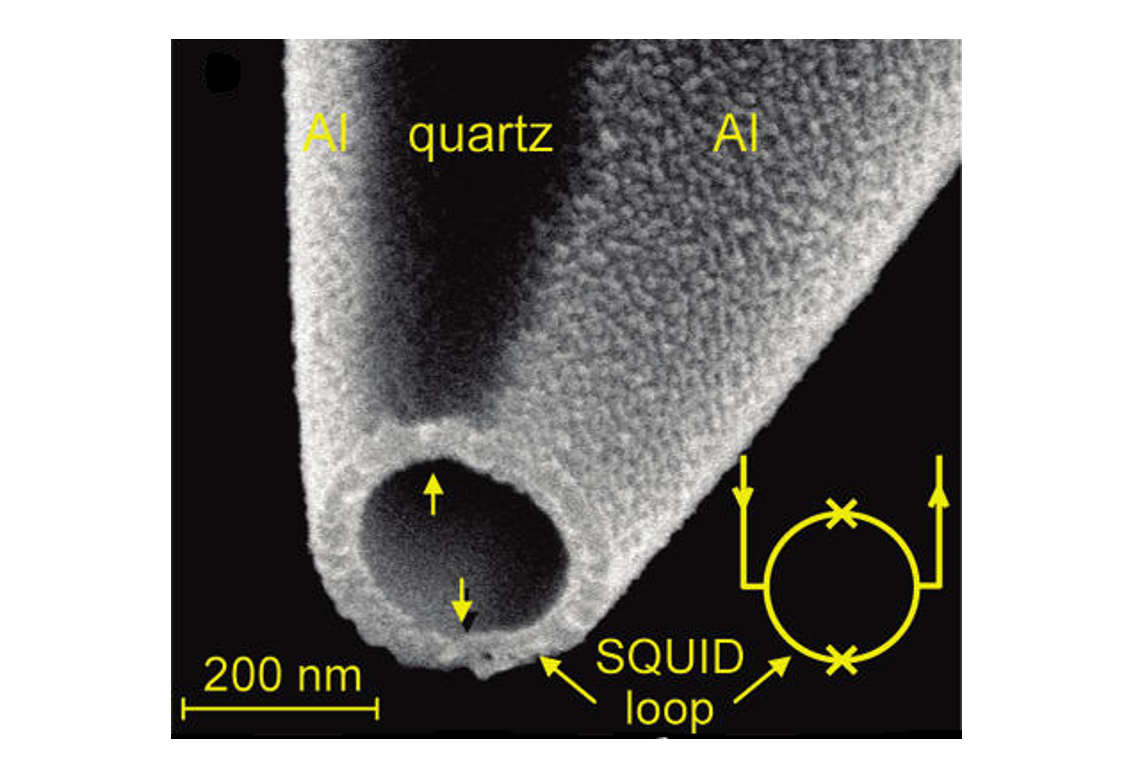

What makes SQUIDs remarkable is their ability to resolve extraordinarily small changes in magnetic fields. This sensitivity arises from quantum effects, which are very responsive to tiny perturbations. As a result, magnetic fields billions of times weaker than the Earth’s ambient field produce detectable signals.
You can use this to measure things like the magnetisation of a sample. The magnetisation tells you about fundamental material properties like currents from electron/atomic motion and particle spins. Perhaps most importantly, magnetometry can tell us about superconductivity, which is critical if we are chasing the holy grail of a room-temperature superconductor.
As an example, SQUIDs can map interesting consequences of superconductivity like the formation and distribution of flux vortices. The picture below shows the flux vortex distribution in YBCO, a family of high-temperature superconducting materials, measured through a technique called scanning SQUID microscopy.
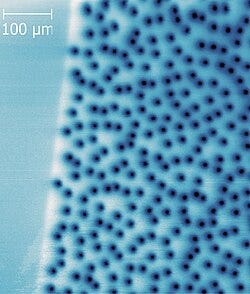

Flux vortices are interesting because superconductors come in two types and flux vortices are a signature of type-II superconductivity. Therefore, vortex distributions can tell you a lot about the fundamental superconductivity properties, like superconducting order parameters.
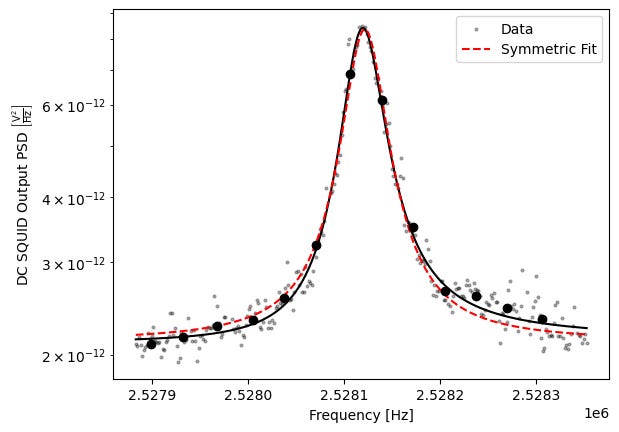
Another example is SQUID noise spectroscopy, which measures tiny magnetic or electrical fluctuations over time. These fluctuations encode information about microscopic degrees of freedom in a material, such as defects and energy loss channels. At a practical level, spectroscopy just produces a continuous time series v(t), a simple signal which is already very neural-net friendly.
As an example of what this can tell us, consider the plot above which shows (Fourier transformed) noise spectroscopy data from my experiment where I use a SQUID to look at noise in a superconducting device. The resulting spectrum has a peak whose width and central frequency encode information about dissipation and energy loss channels in the device materials. In other words, the shape of the noise spectrum directly reflects the microscopic dynamics of the system. These are the types of patterns and relationships neural nets are perfect for.
Since SQUIDs already exist, one might ask whether AI models already use this data. As mentioned earlier, common publicly available materials datasets like the Materials Project largely do not contain the rich experimental data produced in the lab. Instead, lab measurements are first heavily preprocessed by humans into friendly summary tables, which are then fed to today’s models. Of course, some startup may be hiring physicists to bridge this gap, but as far as I know, sensor data is largely untapped.
This is fine as a starting point. The Materials Project contains excellent information about lattice constants and ground-state structures, but who knows what might be hiding in data like the vortex maps of high-Tc materials like YBCO. Could these dynamical patterns hold clues to their unusually high transition temperatures? Neural nets are famously good at finding patterns in troves of data so it’s definitely worth it to unleash them on the data we take in the lab. DeepMind’s GNOME was very successful in predicting new materials, yet its dataset — while rich in simulation-derived quantities like convex hulls — contains no measurements derived from quantum sensors, such as magnetisation maps or noise spectra.
In a theme we will also encounter in biology, current public materials datasets are dominated by static information like lattice constants, while largely missing the dynamic data that quantum sensors naturally provide through techniques such as spectroscopy. While we do have simulation methods to generate dynamic data like ab initio molecular dynamics (AIMD), they are computationally expensive and limited to short timescales (picoseconds). Quantum sensors allow us to measure these dynamics on a longer timescale natively in real-time.
We are almost certainly leaving valuable information on the table by ignoring these fundamental probes, especially if the goal is to understand emergent phenomena like superconductivity. Any serious materials foundation model should draw not only on synthetic simulations, but also on real, sensor-derived data.
Quantum Sensing for Biology
Biology is the other scientific field with lots of promising AI applications. Genomics already produces troves of data which is perfect for AI, but other areas, such as drug discovery, are bottlenecked by the difficulty of generating high-quality experimental data. Fortunately, biology is rich in the type of dynamic signals that can be recorded with quantum sensors. Crucially, as we’ll see, one of the main problems with state-of-the-art protein structure models like AlphaFold is that they are missing exactly this dynamic data.
Brain Perception Models with SQUIDs
SQUIDs are used in biology already, and we have already started seeing cool results when this is paired with AI. Meta trained an AI model to “reconstruct, from brain activity, the images perceived and processed by the brain at each instant”. Naturally, getting high-quality brain activity data is vital for this, and the Meta researchers use a dataset that measures brain activity using a technique called magnetoencephalography (MEG). MEG relies on devices like SQUIDs to measure the tiny magnetic fields produced by currents in neurons. Quantum sensors are needed because they are the only devices with sufficient sensitivity to measure brain neuron activity.
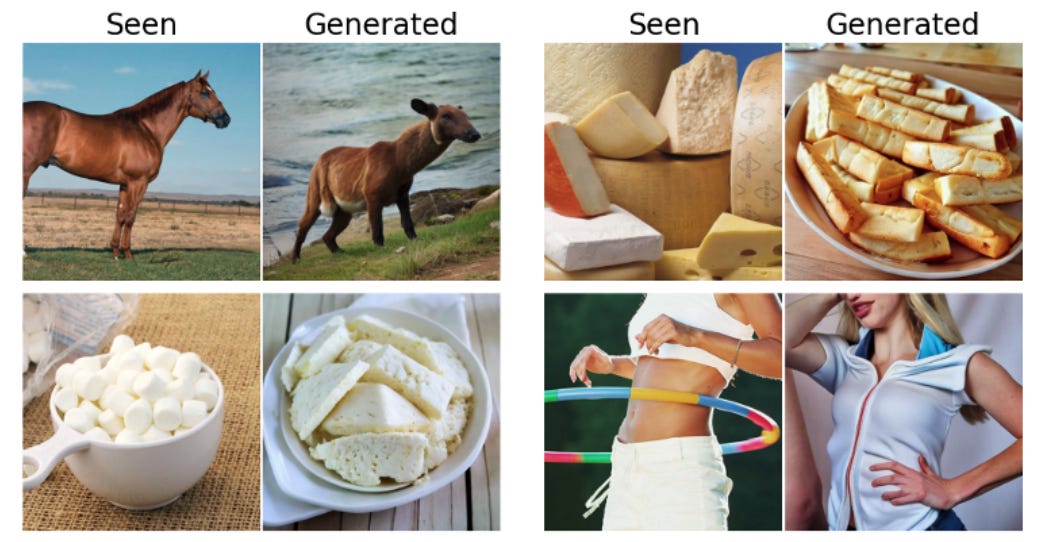

The basic idea is pretty straightforward: show some volunteers images, and then record their brain activity with MEG. Use a portion of that data to train the model by showing it which MEG scans correspond to which images, and then test on the remainder. Some of the results are shown in the picture above. We see that we get pretty good performance — while the image isn’t pixel-for-pixel the same it’s definitely in the right ballpark. It’s quite impressive that we can infer this much already from just brainwaves.
Interestingly, the authors also tried this with fMRI, which is the standard technique for functional imaging of the body. fMRI produces better spatial localisation than MEG, but is fundamentally limited by the timescale of blood flow, so it produces an image once every few seconds. MEG on the other hand is sensitive enough to see neuronal magnetic signals so it can produce thousands of measurements per second. Looking forward to things like brain-computer interfaces, it’s clear that we’ll need to be closer to the MEG end of the spectrum, probably with even better quality quantum sensors to enhance spatial resolution.
Next-Gen Sensors for Cellular Dynamics
SQUIDs are not the only game in town when it comes to sensors, and biology is a wonderful environment to explore this in. One of the most promising next-gen candidates with biological applications are NV-centres, which will give us data at the level of individual cells.
NV-centres are made from diamond, which is a lattice of carbon atoms. An NV-centre is created when one carbon atom is replaced by a nitrogen atom and a neighbouring carbon site is left vacant — hence NV: nitrogen–vacancy. At a basic level, this defect creates a well-behaved quantum system that is extremely sensitive to a whole host of external signals like magnetic fields.
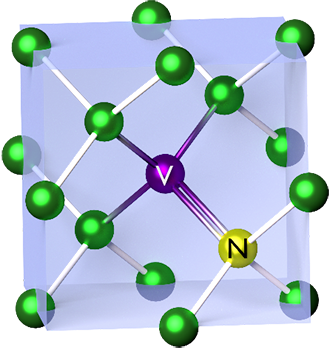
One of the nicest properties of NV-centres is that they work at room temperature. SQUIDs are incredible but they are made of superconductors and so need to be at cryogenic temperatures, which creates huge overhead. NV-centres do not need expensive and bulky cryogenic systems. NV-centres can also be made safe and small enough in “nanodiamond” form to inject into biological tissue, so you can use them for precision sensing at the cell level.
A nice example of using NV-centres for imaging is shown below by a group at Harvard:
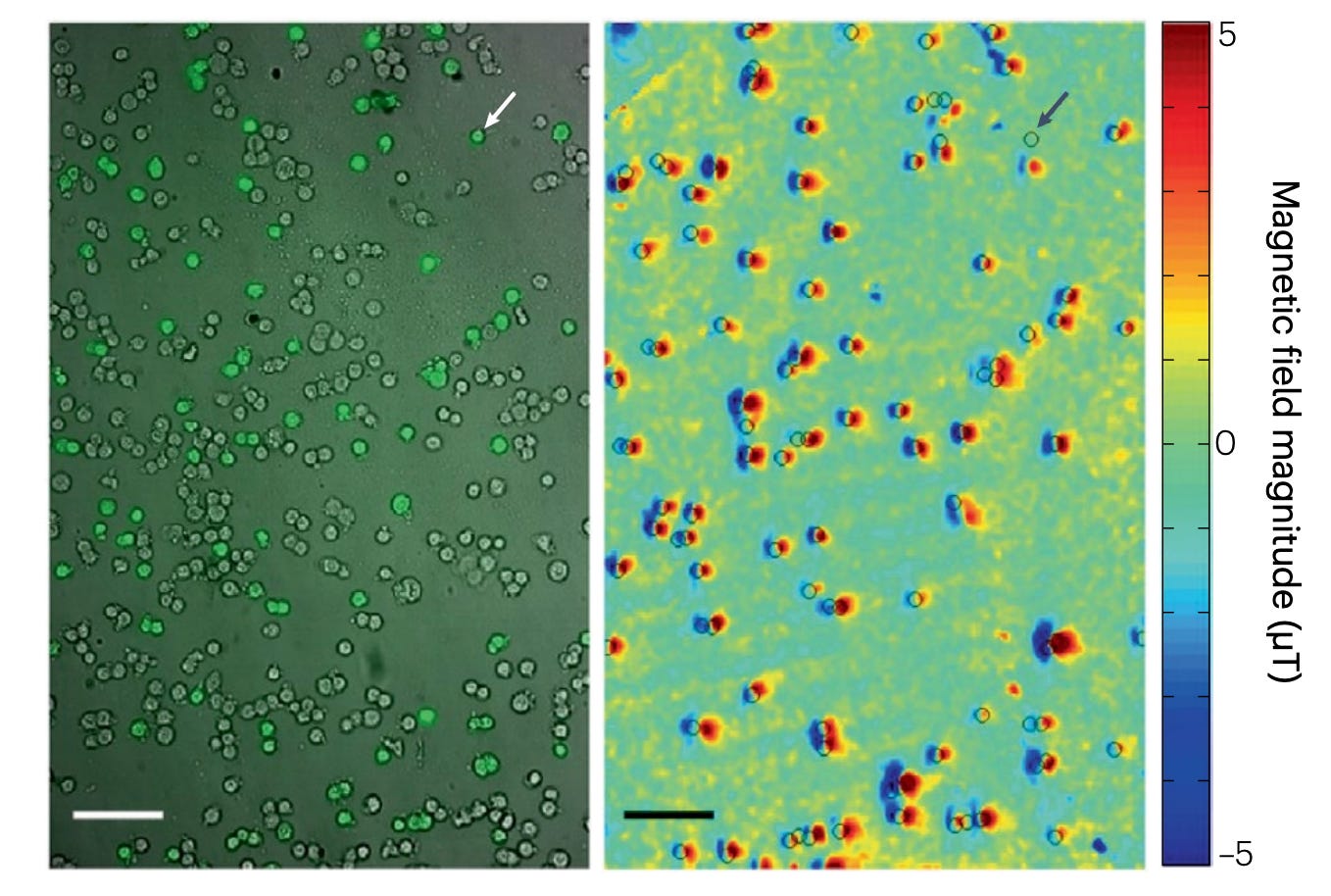

In this experiment, a population of cancer cells is selectively labeled with magnetic nanoparticles (MNPs) which latch onto antibodies targeting a specific biomarker on the cancer cell. These nanoparticles generate tiny local magnetic fields that are detected by a nearby NV-rich diamond sensor, producing a magnetic field map.
Fluorescence imaging — a well-established biological technique — is used in parallel to identify the labeled cells and serve as a ground-truth reference. The strong agreement between the fluorescence signal and the magnetic field map demonstrates that NV-centre magnetometry can reliably detect and spatially resolve biomarker-tagged cells. Such NV-centre techniques are already being commercialised by companies for imaging in medical contexts.
Another critical application of NV centres in biology is spectroscopy. Rather than producing spatial images, NV-centre spectroscopy probes how local magnetic or electric fields fluctuate over time. Much like in SQUID noise spectroscopy for superconducting materials, these fluctuations encode information about underlying dynamics like molecular motion and energy dissipation pathways. Just as the noise spectrum in my own experiment revealed loss channels and microscopic degrees of freedom in a material, NV spectroscopy can reveal how energy flows and redistributes within molecular and cellular systems. This dynamical information is essential for understanding biological function.
Alongside NV-centres, there are plenty of new emerging sensors for biology. A paper from just a few months ago (Aug 2025) demonstrates a quantum sensor based on a fluorescent protein which would be even less invasive in cellular environments than an NV-centre. Meanwhile companies like Precigenetics are working on next generation photonic technology to image cells’ response to drugs in real time, with the ultimate goal of training AI models on that data to enable more efficient drug discovery.
More than for materials, where a lot of the data/techniques already exist, quantum sensors for biology is even more exciting because it’s a far newer field where entire new domains of data are waiting to be created. I’m particularly excited by the ability to map cellular dynamics with greater precision. AlphaFold is no doubt incredible and fully deserves its Nobel Prize, but it’s not perfect. One key reason is that the data from the Protein Data Bank contains information on static structures, and this doesn’t say anything about dynamic behaviour of biomolecules in natural environments like solution. This is exactly the data that quantum sensors can capture.
A Small Word on Quantum Computers
Real-life data is obviously great, but as we now know synthetic simulation data works too. Currently, materials/molecular models are trained on simulations based on density functional theory (DFT) and AIMD. This is fine to start with but it has some limitations. As we saw, AIMD is only short timescale. DFT isn’t good when there are strong electron-electron correlations, as can be the case in high-temperature superconductors. The best way to generate synthetic data for these domains which are fundamentally quantum mechanical would be to run these simulations on a quantum computer. This was very much the original motivation with which Feynman proposed the idea:
Nature isn’t classical, dammit, and if you want to make a simulation of nature, you’d better make it quantum mechanical, and by golly it’s a wonderful problem, because it doesn’t look so easy.
A quantum computer would be gold-standard simulation data for things like quantum many-body emergent phenomena in materials and for modeling molecular interactions in biological contexts. Unfortunately, Feynman was right and it is not easy. The theoretical basis for quantum computing is sound, but the engineering challenges that remain are pretty daunting. This is a potential long-term advantage for Google, because not only does it have world-class AI models, it is perhaps also the world leader in quantum computing at this stage. We are still years to decades away from a working system that can do simulations at the scale required for AI, but this is something to watch for.
As money pours into compute, in many domains the critical bottleneck becomes data. This is especially true for materials and biology. In both of these, we’ve already made some pretty great progress, but the limitations of not having dynamic atomic/molecular level data are increasingly apparent. This is exactly the type of data that quantum sensors can provide today. Quantum computers are a longer term wildcard. They would be the gold-standard for mass producing synthetic data to train on, but it is still unclear how long it will be before we have one good enough to do all the simulations. Overall though, it’s clear that there is a real opportunity here, and unleashing the full potential of quantum could be like an ImageNet moment for materials and molecular models.

Thanks for writing this, it clarifies a lot. How do quantum effects specifically generate new AI data?